Viral Coinfections: Emerging Challenges In Modern Drug Discovery
By Nadine Alvarez, Ph.D., Center for Discovery and Innovation
Viral coinfection is a phenomenon more complex than just the concurrency or sequential infection with multiple viruses in the same host. Previously underreported and now more increasingly recognized, multiple coinfections have been identified across respiratory, enteric, and chronic viral diseases.1–3 When we hear about viral coinfections, we may think that they are always related to the coexistence of two viruses, but this is not always the case. The identification of triple infection has been also documented, especially in seasons where high levels of respiratory viruses circulate and increase hospitalizations. Two recent examples are represented by the detection of respiratory syncytial virus (RSV), human coronavirus OC43, and rhinovirus in a geriatric but immunocompetent patient;4 and the identification of SARS-CoV-2, RSV, and influenza virus in a 52-year-old woman who previously recovered from pneumonia.5
Infants and older adults, immunocompromised individuals, and patients with underlying conditions — such as asthma, chronic obstructive pulmonary disease (COPD), or community-acquired pneumonia (CAP) — represent some of the most vulnerable populations at increased risk of coinfection with viral pathogens.6–8 According to the CDC, 16% of the influenza-associated pediatric deaths during the 2021–2022 influenza season had SARS-CoV-2 coinfection.9 In addition, a previous analysis of the prevalence of mono-infections and coinfections of 13 respiratory viruses over the period of 2013–2018 demonstrated that respiratory viral coinfections were highly predominant in children under five years of age.10 Enteric virus coinfections are commonly identified for all viruses linked to acute gastroenteritis (AGE), including coronavirus, rotavirus, norovirus, astrovirus, adenovirus, and enteroviruses.11 Even when coinfections are more commonly thought of as viruses from different viral families mediating simultaneous infection, previous reports highlight the occurrence of coinfections by species from the same viral genus.12 The extent to which multiple viral infections produce overlapping clinical manifestations or distinct disease outcomes depends largely on whether the interacting pathogens engage in antagonistic or synergistic interactions within the host.
Viral Interplay: How Complex Are Their Interactions?
Viral–viral interactions have been investigated for decades, and a widely recognized classification categorizes these interactions into direct effects, indirect effects, and immunologically mediated influences on viral pathogenicity (Figure 1).13 This phenomenon is extremely intricate, shaped not only by the consequences of coinfections but also by the potential occurrence of superinfection, defined as an infection that occurs after a previous infection, especially when caused by microorganisms that are resistant, or have become resistant, to the antibiotics previously used.14,15 Viral coinfections significantly influence host outcomes, as they can either suppress or enhance viral transmission depending on the nature of the interactions between the co-infecting pathogens.16,17 Different findings have been collected when evaluating the impact of coinfections in viral pathogenicity, with studies reporting no change or even alleviation of the symptoms.18 In other cases, increased effects associated with higher admission rates in intensive care units and development of fibrosis and higher liver-related mortality were seen.19
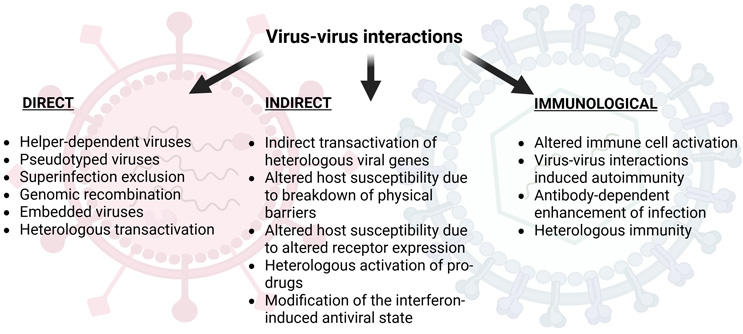
Figure 1. Virus-virus interactions
Collectively, these interactions shape infection dynamics, disease severity, and viral evolution. The bacterial habitat and the interactions with neighboring viruses are two additional key factors that consistently influence patterns of viral coinfection, supported by the fact that host ecology has been identified as a consistent and strong predictor of coinfections across cross-infectivity, culture coinfection, and single-cell coinfection.20 Although evidence supporting the clinical significance of coinfections among different respiratory and enteric viruses remains limited, a well-established association exists between respiratory viral–bacterial coinfections and increased disease severity.21
Rethinking Viral Coinfection Challenges
Coinfections involving multiple pathogenic viruses are of particular concern, as they can exacerbate disease severity and complicate both diagnostic evaluation and therapeutic management. Multiple challenges are related to viral coinfections (Figure 2).
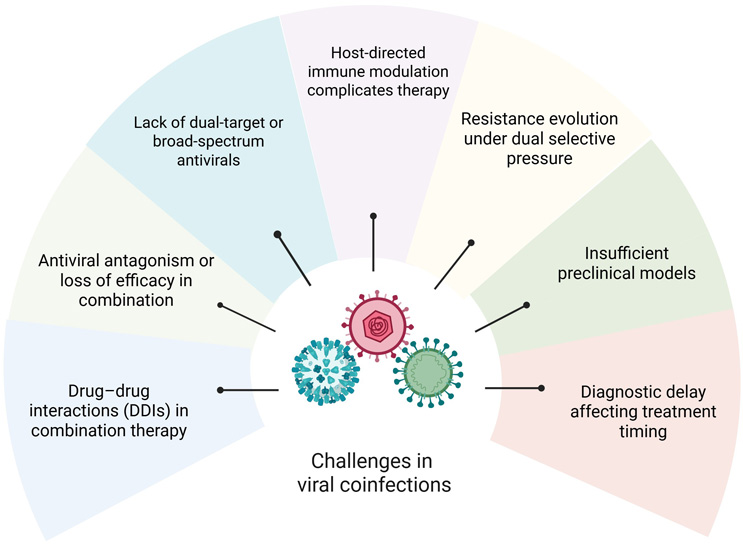
Figure 2. Common challenges in the study of viral coinfections
Although diagnostic approaches have advanced to detect simultaneous viral infections, their sensitivity and specificity are frequently compromised by suboptimal sample quality, sequence contamination, and misinterpretation arising from cross-reactivity among related viral targets.
Therapeutic efficacy in co-administered regimens is especially critical in the context of viral coinfections, where multiple drugs may be required for simultaneous administration. Careful management of drug interactions is crucial, especially in patients receiving anti-retroviral therapy alongside other antiviral agents. However, there is limited guidance for physicians about treatment of viral coinfections. The broad range of possible coinfection combinations complicates the identification of the primary pathogen and makes it difficult to predict their impact on disease severity and clinical outcomes, as different pathogen pairings can influence the course of illness in distinct ways. Despite ongoing efforts to optimize treatment strategies, there remains a pressing need for more effective combination therapies that reduce the occurrence of drug resistance and minimize adverse effects, including those arising from drug toxicity and drug–drug interactions (DDIs).22 On the other hand, dose optimization for drug combinations is often more challenging than for single-drug therapies. Three main challenges are:
- achieving efficient and accurate optimization of drug combinations
- translating findings from laboratory studies to clinical practice
- enabling individualized optimization of combinatorial drug therapies.23
Dose optimization also requires deep pharmacodynamic and pharmacokinetic studies, such as probe drug cocktails, liver microsome and hepatocyte models, in vitro static and dynamic testing, and physiologically based pharmacokinetic models.24
Ideally, broad-spectrum antivirals (capable of inhibiting multiple viruses within the same or close-related families) or pan-antiviral inhibitors (which act across multiple different viral families) should be available for the treatment of coinfections. However, the high mutation rates characteristic of many viruses, particularly RNA viruses, contribute to the rapid emergence of drug resistance. For example, the RNA-dependent RNA polymerase (RdRp) enzyme of SARS‑CoV‑2 exhibits a high frequency of mutations, some of which are unique, that can reduce susceptibility to existing antiviral therapies such as remdesivir and even to certain monoclonal antibodies.25 To mitigate resistance, strategies such as continuous monitoring of viral mutations and the development of therapeutics that target highly conserved viral proteins are suggested.
The success of the therapeutic candidates is often affected by the high variability of the viral surface proteins, which can compromise their antiviral activity. A recent promising strategy to overcome this issue is the development of broad-spectrum antivirals based on small molecules called synthetic carbohydrate receptors, targeting the fact that many viruses have glycans attached to their surface.26 From this study, a couple of molecules demonstrated in vitro potency against six different viruses and significantly reduced SARS-CoV-2 infection in a mouse model. In silico and structural modeling approaches are also widely used tools to advance the identification of potential broad-spectrum antivirals. Two novel peptidomimetics (NIP-22c and CIP-1), reversible covalent inhibitors of SARS-CoV-2 3CLpro, were predicted and further confirmed by in vitro enzymatic and cell-based assays as nanomolar inhibitors of norovirus, enterovirus and rhinovirus.27
Experimental Tools And Strategic Approaches To Address Viral Coinfections Challenges
Different platforms are currently in use for the exploration and understanding of the viral coinfections dynamics, including in vitro 2D models using immortalized cell lines; air-liquid interface (ALI) cultures; ex vivo 3D systems, such as organoids and lung-on-a-chip platforms; iPSC-derived models; precision-cut lung slices (PCLS); and in vivo animal models.28 While each platform has inherent limitations, such as incomplete representation of all epithelial cell types, genetic variability, limited assay duration, or organoid viability, each also offers distinct advantages that collectively help address knowledge gaps in the study of viral coinfections. Nevertheless, the development of novel experimental models remains a priority. Alternative methods include the use of human lung buds derived from human embryonic stem cells (hESCs) and cultured on micropatterned substrates for high-throughput identification of potential novel therapeutics.29 In addition, mathematical and computational models that incorporate multiple viruses are used to simulate dynamic infection, predict time courses, and estimate peak viral loads through quantitative analyses.30 However, these predictive models require validation in physiologically relevant in vivo contexts to accurately capture immune responses and overcome the limitations of in silico predictions. The integration of artificial intelligence (AI), machine learning, and deep learning tools to study clinical virology plays a critical role in the enhancement of diagnostic precision, therapeutic interventions, epidemiological monitoring, processing of genomic data, and analysis of microscopic imaging.31
Advancing the understanding of the molecular mechanisms and clinical consequences of viral coinfections requires the integration of multi-omics approaches, including genomics, transcriptomics, proteomics, and metabolomics, alongside the development and rigorous characterization of experimental models that more accurately recapitulate human physiology. Progress in this field will depend on coordinated efforts spanning in vitro and ex vivo systems, preclinical models, and well-designed clinical studies that faithfully reflect the complexity of human pathology and disease manifestations. However, many of these platforms are resource-intensive, demand specialized technical expertise, and often require access to biosafety level 3 (BSL-3) facilities, based on the pathogen addressed. Securing sustained long-term funding also remains a significant barrier, particularly for extended studies. In parallel, comprehensive longitudinal cohorts and robust surveillance programs are indispensable for defining the long-term health consequences of viral coinfections, their impact on disease progression, and their influence on future susceptibility and risk trajectories. Collaborative efforts that bring together multidisciplinary academic teams, clinicians, and pharmaceutical partners are critical to advancing promising preclinical candidates toward rigorous validation and IND enabling studies, as well as to translating laboratory discoveries into more effective prevention strategies, diagnostic tools, and therapeutic interventions.
References
- Yan X., Li K., Lei Z., Luo J., Wang Q., Wei S. Prevalence and associated outcomes of coinfection between SARS-CoV-2 and influenza: A systematic review and meta-analysis. Int. J. Infect. Dis. 2023;136:29–36. doi: 10.1016/j.ijid.2023.08.021.
- Halabi K.C., Wang H., Leber A.L., Sánchez P.J., Ramilo O., Mejias A. Respiratory syncytial virus and SARS-CoV-2 coinfections in children. Pediatr. Pulmonol. 2022;57:3158–3160. doi: 10.1002/ppul.26127.
- Tang M.L., Li Y.Q., Chen X., Lin H., Jiang Z.C., Gu D.L., Chen X., Tang C.X., Xie Z.Q. Co-Infection with Common Respiratory Pathogens and SARS-CoV-2 in Patients with COVID-19 Pneumonia and Laboratory Biochemistry Findings: A Retrospective Cross-Sectional Study of 78 Patients from a Single Center in China. Med. Sci. Monit. 2021;27:e929783. doi: 10.12659/MSM.929783.
- Davido B, Jaffal K, Gault E, Bourlet S, Beaune S. Back to the future of viruses: a case of triple coinfection caused by respiratory syncytial virus, human coronavirus OC43, and rhinovirus. Int J Infect Dis. 2023;130:205-207. doi: 10.1016/j.ijid.2023.03.024.
- Guido G, Lalle E, Mosti S, Mencarini P, Lapa D, Libertone R, Ianniello S, Ricciuto GM, Vaia F, Maggi F, Palmieri F. Recovery from Triple Infection with SARS-CoV-2, RSV and Influenza virus: A case report. J Infect Public Health. 2023;16(7):1045-1047. doi: 10.1016/j.jiph.2023.05.001.
- Wiseman DJ, Thwaites RS, Ritchie AI, Finney L, Macleod M, Kamal F, Shahbakhti H, van Smoorenburg LH, Kerstjens HAM, Wildenbeest J, Öner D, Aerssens J, Berbers G, Schepp R, Uruchurtu A, Ditz B, Bont L, Allinson JP, van den Berge M, Donaldson GC, Openshaw PJM, Wedzicha J; RESCEU Investigators. Respiratory Syncytial Virus-related Community Chronic Obstructive Pulmonary Disease Exacerbations and Novel Diagnostics: A Binational Prospective Cohort Study. Am J Respir Crit Care Med. 2024;210(8):994-1001. doi: 10.1164/rccm.202308-1320OC.
- Santus P, Danzo F, Signorello JC, Rizzo A, Gori A, Antinori S, Gismondo MR, Brambilla AM, Contoli M, Rizzardini G, Radovanovic D. Burden and Risk Factors for Coinfections in Patients with a Viral Respiratory Tract Infection. Pathogens. 2024;13(11):993. doi: 10.3390/pathogens13110993.
- Radovanovic D, Sotgiu G, Jankovic M, Mahesh PA, Marcos PJ, Abdalla MI, Di Pasquale MF, Gramegna A, Terraneo S, Blasi F, Santus P, Aliberti S, Reyes LF, Restrepo MI; GLIMP Study Group. An international perspective on hospitalized patients with viral community-acquired pneumonia. Eur J Intern Med. 2019;60:54-70. doi: 10.1016/j.ejim.2018.10.020.
- Adams K, Tastad KJ, Huang S, Ujamaa D, Kniss K, Cummings C, Reingold A, Roland J, Austin E, Kawasaki B, Meek J, Yousey-Hindes K, Anderson EJ, Openo KP, Reeg L, Leegwater L, McMahon M, Bye E, Poblete M, Landis Z, Spina NL, Engesser K, Bennett NM, Gaitan MA, Shiltz E, Moran N, Sutton M, Abdullah N, Schaffner W, Talbot HK, Olsen K, Staten H, Taylor CA, Havers FP, Reed C, Budd A, Garg S, O'Halloran A, Brammer L. Prevalence of SARS-CoV-2 and Influenza Coinfection and Clinical Characteristics Among Children and Adolescents Aged <18 Years Who Were Hospitalized or Died with Influenza-United States, 2021-22 Influenza Season. MMWR Morb Mortal Wkly Rep 2022;71:1589–1596. doi: 10.15585/mmwr.mm7150a4.
- Mandelia Y., Procop G.W., Richter S.S., Worley S., Liu W., Esper F. Dynamics and predisposition of respiratory viral co-infections in children and adults. Clin. Microbiol. Infect. 2021;27:e1–e631. doi: 10.1016/j.cmi.2020.05.042.
- Holtz L.R., Cao S., Zhao G., Bauer I.K., Denno D.M., Klein E.J., Wang D. Geographic variation in the eukaryotic virome of human diarrhea. Virology. 2014;468–470:556–564. doi: 10.1016/j.virol.2014.09.012.
- Ruggeri F.M., Delogu R., Petouchoff T., Tcheremenskaia O., Petris S.D., Fiore L. RotaNet-Italy study group molecular characterization of rotavirus strains from children with diarrhea in Italy, 2007–2009. J. Med. Virol. 2011;83:1657–1668. doi: 10.1002/jmv.22163.
- DaPalma T, Doonan BP, Trager NM, Kasman LM. A systematic approach to virus-virus interactions. Virus Res. 2010;149(1):1-9. doi: 10.1016/j.virusres.2010.01.002.
- Musuuza JS, Watson L, Parmasad V, Putman-Buehler N, Christensen L, Safdar N. Prevalence and outcomes of co-infection and superinfection with SARS-CoV-2 and other pathogens: A systematic review and meta-analysis. PLoS One. 2021;16(5):e0251170. doi: 10.1371/journal.pone.0251170.
- Antibiotic resistance threats in the United States, 2013. U.S Department of Health and Human Services. Centers for Disease Control and Prevention https://www.cdc.gov/drugresistance/pdf/ar-threats-2013-508.pdf. Accessed on March 4, 2026.
- Vasilakis N, Holmes EC, Fokam EB, Faye O, Diallo M, Sall AA, Weaver SC. Evolutionary processes among sylvatic dengue type 2 viruses. J Virol. 2007;81(17):9591-5. doi: 10.1128/JVI.02776-06.
- Kent RJ, Crabtree MB, Miller BR. Transmission of West Nile virus by Culex quinquefasciatus say infected with Culex Flavivirus Izabal. PLoS Negl Trop Dis. 2010;4(5):e671. doi: 10.1371/journal.pntd.0000671.
- Guan Z, Chen C, Li Y, Yan D, Zhang X, Jiang D, Yang S, Li L. Impact of Coinfection With SARS-CoV-2 and Influenza on Disease Severity: A Systematic Review and Meta-Analysis. Front Public Health. 2021;9:773130. doi: 10.3389/fpubh.2021.773130.
- Amin J, Law MG, Bartlett M, Kaldor JM, Dore GJ. Causes of death after diagnosis of hepatitis B or hepatitis C infection: a large community-based linkage study. Lancet. 2006;368(9539):938-45. doi: 10.1016/S0140-6736(06)69374-4.
- Díaz-Muñoz SL. Viral coinfection is shaped by host ecology and virus-virus interactions across diverse microbial taxa and environments. Virus Evol. 2017;3(1):vex011. doi: 10.1093/ve/vex011.
- Azevedo M, Mullis L, Agnihothram S. Viral and Bacterial Co-Infection and Its Implications. SciFed Virol Res J. 2017;1(1):10.23959/sfjv-1000002. doi: 10.23959/sfjv-1000002.
- Glezen WP, Taber LH, Frank AL, Kasel JA. Risk of primary infection and reinfection with respiratory syncytial virus. Am J Dis Child. 1986;140:543–546. doi: 10.1001/archpedi.1986.02140200053026.
- Wang B, Warden AR, Ding X. The optimization of combinatorial drug therapies: Strategies and laboratorial platforms. Drug Discov Today. 2021 Nov;26(11):2646-2659. doi: 10.1016/j.drudis.2021.07.023.
- Sun L, Mi K, Hou Y, Hui T, Zhang L, Tao Y, Liu Z, Huang L. Pharmacokinetic and Pharmacodynamic Drug-Drug Interactions: Research Methods and Applications. Metabolites. 2023;13(8):897. doi: 10.3390/metabo13080897.
- Batool S, Chokkakula S, Jeong JH, Baek YH, Song MS. SARS-CoV-2 drug resistance and therapeutic approaches. Heliyon. 2025 Jan 15;11(2):e41980. doi: 10.1016/j.heliyon.2025.e41980. PMID: 39897928; PMCID: PMC11786845.
- Ezzatpour S, Thakur K, Erzoah Ndede K, Buchholz DW, Choi A, Imbiakha B, Carter J, Onofrei D, Eaton B, Postnikova E, Murphy M, Tapia BC, Bello D, Pasari S, Russo A, Babayev M, Holland GP, Holbrook MR, Caddy SL, Moran SJ, Davachi SM, Monreal IA, Sahler J, Ortega V, Miranda JM, Whittaker GR, Jager MC, Bhagwat SK, Chopra P, Jan-Boons G, Marianski M, Braunschweig AB, Aguilar HC. Broad-spectrum synthetic carbohydrate receptors (SCRs) inhibit viral entry across multiple virus families. Sci Adv. 2025;11(35):eady3554. doi: 10.1126/sciadv.ady3554.
- Patel D, De R, Azadi N, Lee S, Shooter S, Amichai S, Zhou S, Monroe D, Mahanke C, McBrayer TR, Muczynski M, Al-Homoudi A, Engel J, Bochkov YA, Gern JE, Kovari LC, Amblard F, Schinazi RF. Discovery of broad-spectrum antivirals targeting viral proteases using in silico structural modeling and cellular analysis. Antiviral Res. 2025;241:106245. doi: 10.1016/j.antiviral.2025.106245.
- Yazici O, Vanetti C, Clerici M, Biasin M. Experimental Models to Investigate Viral and Cellular Dynamics in Respiratory Viral Co-Infections. Microorganisms. 2025;13(11):2444. doi: 10.3390/microorganisms13112444.
- Rosado-Olivieri EA, Razooky B, Le Pen J, De Santis R, Barrows D, Sabry Z, Hoffmann HH, Park J, Carroll TS, Poirier JT, Rice CM, Brivanlou AH. Organotypic human lung bud microarrays identify BMP-dependent SARS-CoV-2 infection in lung cells. Stem Cell Reports. 2023;18(5):1107-1122. doi: 10.1016/j.stemcr.2023.03.015.
- Srivastava S, Dobrovolny HM. Quantitative analysis of respiratory viral triple infections: Examining within host dynamics of Influenza, RSV, and SARS-CoV-2. Math Biosci Eng. 2025;22(11):2852-2869. doi: 10.3934/mbe.2025105.
- Padhi A, Agarwal A, Saxena SK, Katoch CDS. Transforming clinical virology with AI, machine learning and deep learning: a comprehensive review and outlook. Virusdisease. 2023 Sep;34(3):345-355. doi: 10.1007/s13337-023-00841-y.
About The Author
 Nadine Alvarez, Ph.D., serves as a supervisor research assistant member, infectious diseases at the Center for Discovery and Innovation (CDI), where she leads projects focused on the discovery and advancement of novel therapeutic candidates for antiviral and antibacterial infections.
Nadine Alvarez, Ph.D., serves as a supervisor research assistant member, infectious diseases at the Center for Discovery and Innovation (CDI), where she leads projects focused on the discovery and advancement of novel therapeutic candidates for antiviral and antibacterial infections.
She earned her bachelor’s degree in biology, a Master of Science in biochemistry with a specialization in molecular biology, and a Ph.D. in health sciences, all from the University of Havana, Cuba. She subsequently transitioned into drug discovery at the Public Health Institute of Research at Rutgers University. Over the past five years at CDI, her work has focused on the identification and development of small molecules as innovative treatment options for viral infections. More recently, her research has centered on Mpox.
