The Missing Data Set In Drug Discovery: Patient Experience As Decision Data
By Elisha Lowe, RN, BSN, MBA

I have worked across clinical care and the life sciences industry: as an ICU nurse; as a patient who has moved through the same system I spent my career working in; and across commercial and medical affairs roles in medtech, diagnostics, and specialty pharmacy. In each of those settings, I have watched the same gap appear: the distance between how a therapy is designed and how it is actually experienced by the people it is meant to help. By the time that gap becomes visible in a trial, the decisions that created it have already been made.
Drug Discovery Operates On An Expanding Biological Data Infrastructure
Drug discovery runs on biological data: genomic data sets, biomarker discovery platforms, multi-omics analysis, and machine learning-assisted target identification. Yet development programs continue to encounter recurring barriers once therapies reach human trials. Recruitment falls short, and endpoints fail to reflect outcomes that patients consider meaningful. Treatment regimens prove difficult to sustain. Some of those failures are biological, while others emerge when therapies reach real patients and collide with how disease is lived.
This piece does not argue that patient experience data solves the biology. It argues there are specific R&D decision points where structured patient experience variables could inform decisions that are currently made without them.
The Dominant Data Model In Discovery
The discovery enterprise is optimized around biological data sets: genomic associations, pathway mapping, target validation, and preclinical disease models. These signals are powerful because they are quantitative, structured, and computationally compatible. They have made possible a generation of targeted therapies that would have been unimaginable 30 years ago.
But biological data sets primarily describe disease at the molecular level, not how it is experienced in daily life: the symptom burden a patient manages between clinic visits, the functional limitations that shape whether they can work or care for their family, the tolerability thresholds that determine whether they continue treatment, and the behavioral constraints that affect adherence in ways no pharmacokinetic model predicts.
And they are almost entirely absent from early discovery data sets.1
Translational Failure Often Reveals The Gap
Despite improvements in target identification and molecular screening, roughly 90% of drug candidates entering clinical trials fail. Many of these failures reflect genuine biological complexity. The science was right, but the biology did not cooperate. Others arise from a different kind of mismatch. The proportion attributable to each is not well quantified, but the pattern is observable across multiple programs.
Consider sarcoidosis, a systemic inflammatory disease affecting multiple organ systems. Patients consistently report fatigue, cognitive impairment, and the inability to predict functional capacity from day to day as their most disabling symptoms. These are not captured by standard inflammatory biomarkers. A therapy that successfully suppresses granuloma formation while leaving fatigue and cognitive fog unaddressed may register as biologically active and still fail to meaningfully improve patient lives or fail to demonstrate benefit on patient-centered endpoints in a trial. In some cases, these symptoms may not be mechanistically linked to the pathway being targeted, which makes the disconnect even more important to identify early.
For chronic diseases broadly, patients prioritize improvements in fatigue, mobility, cognitive function, and daily independence. These priorities are rarely visible during early discovery stages. By the time they surface in clinical trials, the therapy's molecular architecture has already been established, and the development timeline has been set.
Patient Experience Is Already Used, But Too Late
The industry has made real progress on patient perspectives. Patient-reported outcomes (PRO), natural history studies, and the FDA’s Patient-Focused Drug Development (PFDD) programs have moved patient voices into the development process. These are real advances.
But they are primarily instruments for evaluating therapies, not for shaping early discovery decisions. In most development programs, patient experience data enters the process after target selection and lead optimization are complete. The molecular architecture has been committed. The delivery mechanism has been chosen. By the time a PRO instrument is designed or a PFDD meeting convenes, the fundamental decisions about what the drug is and how it works have already been made.1,2
Discovery Decision Points Where Patient Experience Data Matters
Several discovery milestones represent meaningful opportunities to integrate patient experience data before molecular commitments are made. The table below maps these decision points against the biological data currently used and the patient experience variables that could complement them.
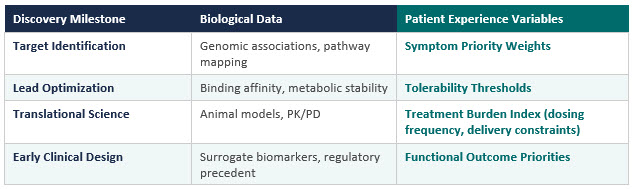
These are not replacements for biological inputs. They are parallel signals that address questions biological data alone does not fully answer: what do patients need this therapy to do for their daily lives, and what constraints will determine whether they use it?
Symptom priority weights: a structured ranking of which symptoms most affect daily function, derived from structured coding of patient narratives and expressed as a weighted distribution across a population — not always captured in standard clinical measures, but consistently reported as most limiting by patients.
Tolerability thresholds: the point at which a specific side effect or burden causes a patient to discontinue, quantified as discontinuation probability associated with specific side effects.
Treatment burden index: a composite of the time, logistics, physical effort, and dependency required to maintain a therapy — distinct from clinical tolerability and rarely captured in trial protocols.
Functional outcome priorities: what getting better actually looks like in a patient's daily life, translated into ranked functional endpoints that can inform endpoint selection and success criteria.
Consider a chronic disease program where candidates are comparable on key biological and pharmacological dimensions and both an oral therapy and an infused biologic are scientifically viable. If efficacy differences are modest, the decision often comes down to PK, manufacturability, and competitive precedent. Patient experience data on treatment burden, logistics, and daily life disruption could inform that calculus. A patient population that depends on public transportation, works rotating shifts, or manages caregiving responsibilities does not have equal access to infusion center visits. That is a real constraint that affects long-term adherence, and adherence determines real-world efficacy. Integrating that information at the point of modality selection, before chemistry is committed, is different from discovering it retrospectively in a Phase 3 dropout analysis, when the cost of change is already prohibitive.
Patient experience data is not a universal input. Target identification driven by genetic association or structural biology does not require patient narrative input to establish mechanism. Safety margins, pharmacokinetic constraints, and tissue accessibility are not negotiable based on patient preference. Where candidates are comparable on key biological and pharmacological dimensions, patient experience variables can inform the decision. Where they are not comparable, biology governs. The purpose of integrating patient experience data is not to override mechanism. It is to prevent programs from being optimized around outcomes that are measurable but not meaningful.
Converting Experience Into Data And Expanding The Discovery Data Environment
Biological information is inherently structured. A binding affinity measurement, a genomic variant call, a pharmacokinetic curve: these are already in formats that computational systems can analyze. Patient experience data is not. It lives in interviews, patient forums, advisory testimony, support groups, and qualitative research reports. Historically, this information has been treated as anecdotal rather than analytical. Useful for context, inadequate for evidence.
Structured qualitative coding methodologies and computational text analysis now make it possible to extract repeatable, machine-readable signals from patient narratives, converting what was once treated as anecdote into analyzable evidence.3,4 Research has consistently found, however, that natural language processing (NLP) outputs require domain expert validation to translate into clinically and operationally meaningful insights.4
The methodological infrastructure is sufficiently developed to support this work. What the field has lacked is agreement that it belongs in discovery.
Drug discovery has steadily expanded the range of biological datasets it uses to understand disease. Genomics gave way to proteomics. Proteomics to multi-omics. Each expansion increased the field's ability to characterize what is happening inside the body. The next expansion may not be deeper into biology but broader in what we are willing to treat as data.
Conclusion
Drug discovery has become highly effective at capturing molecular detail. The tools available to modern scientists, for target identification, for lead optimization, for translational modeling, are more powerful than anything the field has had before.
Yet the experience of disease, the functional burden, the treatment trade-offs, the daily constraints patients live with, remains difficult to incorporate into early scientific decisions — not because it is unknowable, but because it has not yet been treated as data.
Closing this gap does not replace biological discovery. It expands the data environment in which discovery operates. The therapies most likely to survive the translational journey are probably not just the ones that are most biologically precise. They are the ones whose design was informed, from the beginning, by what it actually means to live with the disease they are trying to treat.
References
- Almeida D, Umuhire D, Gonzalez-Quevedo R, et al. Leveraging patient experience data to guide medicines development, regulation, access decisions and clinical care in the EU. Front Med (Lausanne). 2024;11:1408636. doi:10.3389/fmed.2024.1408636
- U.S. Food and Drug Administration. Patient-Focused Drug Development: Collecting Comprehensive and Representative Input. Guidance for Industry, Food and Drug Administration Staff, and Other Stakeholders. 2020. Available at: https://www.fda.gov/regulatory-information/search-fda-guidance-documents/patient-focused-drug-development-collecting-comprehensive-and-representative-input
- Sim JA, Huang X, Horan MR, Stewart CM, Robison LL, Hudson MM, Baker JN, Huang IC. Natural language processing with machine learning methods to analyze unstructured patient-reported outcomes derived from electronic health records: a systematic review. Artif Intell Med. 2023;146:102701. doi:10.1016/j.artmed.2023.102701
- Feizollah A, Lin CY, O'Malley L, Thompson W, Listl S, Byrne M. The use of natural language processing to interpret unstructured patient feedback on health services: scoping review. J Med Internet Res. 2025;27:e72853. doi:10.2196/72853
About The Author
 Elisha Lowe, RN, BSN, MBA, is founding principal of IRL Life Sciences Partners and founder of Patient Meets Science, a platform connecting chronically ill patients with life sciences leaders to connect patient experience with commercial strategy. A healthcare and life sciences veteran with 25+ years of experience, she has worked alongside commercial and medical affairs teams across specialty pharmacy, diagnostics, medtech, and patient access programs. Her clinical background spans ICU nursing and rare disease care.
Elisha Lowe, RN, BSN, MBA, is founding principal of IRL Life Sciences Partners and founder of Patient Meets Science, a platform connecting chronically ill patients with life sciences leaders to connect patient experience with commercial strategy. A healthcare and life sciences veteran with 25+ years of experience, she has worked alongside commercial and medical affairs teams across specialty pharmacy, diagnostics, medtech, and patient access programs. Her clinical background spans ICU nursing and rare disease care.
